Download My CV
| ajit_biosketch-som.docx | |
| File Size: | 52 kb |
| File Type: | docx |
Personal statement
I gained extensive research experience in the field of structural biology and biochemistry during my Ph.D. dissertation work. After obtaining my Ph.D. I was interested in applying biophysical techniques to medically relevant problems, and with that intention, I went to the University of Cambridge for postdoctoral training. To conduct independent research, I moved to India and joined Lovely Professional University as an assistant professor. Being a private university there was very limited research opportunity, so I decided to obtain more training in biophysics and gain expertise in cell biology. With that motivation, I joined Sharon Campbell’s lab at UNC Chapel Hill. Two projects which had direct applications to human health interested me the most. The first project was to characterize the function of Gαi protein in intracellular pH sensing, which might be highly relevant to ischemic heart diseases. Using several biophysical techniques and cell-based assay, I identified a novel mechanism of pH sensing by Gai and now desire to expand these studies disease relevant system. The second project is cancer drug discovery by targeting KRAS protein and I identified a novel small molecule that inhibits KRAS protein. My future research involved further characterization of this molecule to treat cancer.
Research work
|
Postdoctoral research at UNC, USA
In 2018 I moved to Sharon Campbell’s Lab at UNC-CH for my second postdoctoral training. When I joined Sharon’s Lab, we discussed several unanswered research questions and I became particularly interested in investigating the potential role of G-protein in pH sensing and regulation. Answering this question would help us understand the basic mechanism of pH sensing in several diseases where cellular pH gets altered. To characterize the pH sensing ability of Gαi, I optimized the high yield of Gαi purification from E.Coli. Later I learned Circular dichroism and performed CD for Gαi at different pH to show that pH indeed affects the stability of Gαi. Later I performed NMR to show that pH affects the dynamic property of Gαi. Later to identify the pH sensing residues in Gαi, I generated more than 15 mutants proteins and tested their pH sensing ability. This extensive screening helped me to discover the key residues of Gαi involved in pH sensing. Furthermore, to understand the mechanism of pH sensing by these residues I learned molecular dynamics (MD) and performed MD for Gαi at different pH to show that lower pH results in protonation of the identified charged resides which breaks the electrostatic interaction of charged residues and leads to order to disorder transition of protein. To understand the biological significance of the pH sensing ability of Gαi, I trained in multiple cell biology techniques. I was then able to optimize several methods to alter and monitor the intracellular pH. I showed a good correlation in the change in intracellular pH by changing extracellular pH or treating the cell with FCCP drug or changing the percentage supply of CO2 to the cell. I also perform a HEK293 cell-based BRET assay to show that the interaction of Gαi with Gβγ and their respective activation might be pH-dependent. We further expanded this study in another isoform of Gαi and showed a potentially conserved mechanism of pH sensing by different isoforms of Gαi. My work to characterize a novel function of G-protein as a pH sensor and modulator is incorporated in a manuscript currently in preparation. I am also involved in two other projects one in collaboration with Dr. Aube Jeff, UNC-CH, and the other with Mirati Therapeutics. In these projects, we are trying to identify novel KRAS inhibitors to treat cancer and also understating the mechanism of action of MRTX849 (discovered by Mirati Therapeutics) which is in phase II clinical trial. I have now identified a novel small molecule (UNC6126) that binds to KRAS with micromolar affinity. I used HADDOCK docking to uncover the binding mode UNC6126 with KRAS and the manuscript for this work is under preparation. |
|
Postdoctoral research at University of Cambridge, UK
Right after my Ph.D., I joined Dr. Louise Boyle's lab as a postdoctoral fellow in the Department of Pathology, at the University of Cambridge in July 2016. During my postdoctoral studies, I studied the regulation of peptide loading on major histocompatibility complex (MHC I) by TAPBPR protein to understand the nonclassical peptide presentation pathway. MHC I is a very difficult protein to purify and I optimized the refolding of MHC protein and also the expression and purification of TAPBPR protein from HEK-293F mammalian cells. I tried to make a complex of MHC and TAPBPR by several strategies to get the crystal for structural studies. I also modified the Piggy Expression vector by molecular cloning technique and used that modified vector to make a stable cell line for TAPBPR expression in HEK293 cells. |
|
As an assistant professor at Lovely Professional University, India Right after my first postdoc at the University of Cambridge, UK, I moved to my home country India and joined Lovely Professional University (the largest private university in India) and taught undergraduate and master's students there. The subjects which I thought there were Biochemistry, Cell Biology, Basic Biology and Molecular Biology I also guided four students in their master’s dissertation research. To get more research experience, I moved to the USA for my second postdoctoral study at Sharon Campbell Lab. I still collaborate with my students from Lovely Professional University and recently published multiple papers (also as a corresponding author). |
|
Ph.D. thesis work
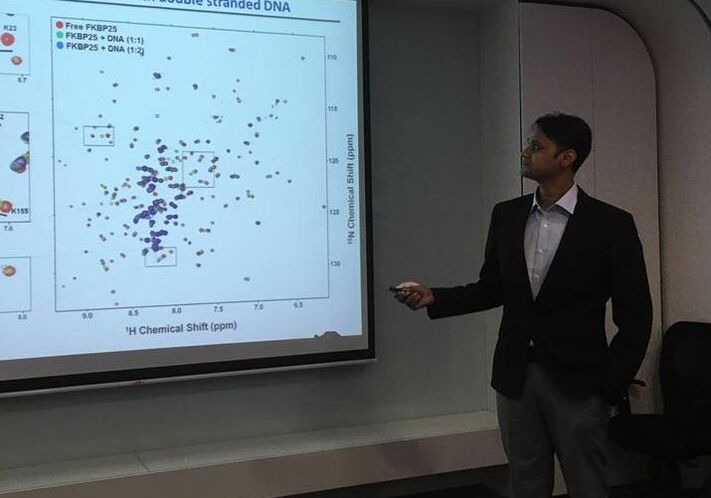
During the first semester of my Ph.D., I worked with Prof. Ravi Kambadur and I tried to elucidate the mechanism of insulin sensitivity in myostatin knockout mice fed on a high-fat diet. During this period, I extensively performed RT-PCR, cell culture work, western blotting and mouse work. I also completed an animal handling course in Singapore. Then I joined Prof. Yoon Ho Sup's lab and finished my Ph.D. under his supervision. During my Ph.D., I cloned, expressed, and purified the human FKBP25 protein in E.coli. Purified protein was further used to determine the NMR solution structure of FKBP25. I also solved the crystal structure of FKBD25 in complex with the immunosuppressive drug FK506 at a resolution of 1.8 Å. Later using NMR, ITC and EMSA, I showed that FKBP25 binds with dsDNA in a sequence-independent manner. We generated several mutants of FKBP25 to abolish FKBP25-DNA interaction. Further, we could map the DNA binding site of FKBP25. We performed HADDOCK docking to get the FKBP25-DNA complex model. Later we also showed that FKBP25 forms a ternary complex with DNA and YY1 (a transcription factor) and this ternary complex formation enhance the transcriptional repression activity of YY1. We also tried to crystallize the FKBP25-DNA complex to solve the crystal structure of the complex. In collaboration with Dr. Phan, we showed that FKBP25 had the potential to bind to G-quadruplex DNA. I published three high-impact first-author papers during my Ph.D. research. |
|
Pre-PhD research: I have done six months M.Sc. dissertations under the supervision of Prof. Hussain Muanavar, MKU, India. I was employed in cloning a nonsense suppressor tRNA named SupE44. I cloned this suppressor tRNA and reported its multi-copy effect in E.coli. During my first year of my Master, I did three months summer internship at the Indian Institute of Science Education and Research (IISER) Bhopal, India under Prof. Vikas Jain. During the project, we isolated 8 different metal-resistant bacteria from the Bhopal gas tragedy site. I identified and characterized all the bacteria and also determined MIC (minimal inhibitory concentration) under different metal stress conditions using Co, Mg, Cu, Ag, Hg, Cr etc. |
Skills
X-ray crystallography
Western Blotting
Fluorescence Spectrometer
Protein purification
Mutagenesis
Cell culture
PCR
NMR
ITC
Western Blotting
Fluorescence Spectrometer
Protein purification
Mutagenesis
Cell culture
PCR
NMR
ITC